PointNet is a deep learning architecture for classification and segmentation of 3D point clouds. As stated by Qi et al (Qi et al., 2017), the PointNet is a
[…] novel deep net architecture that consumes raw point cloud (set of points) without voxelization or rendering. It is a unified architecture that learns both global and local point features, providing a simple, efficient and effective approach for a number of 3D recognition tasks.
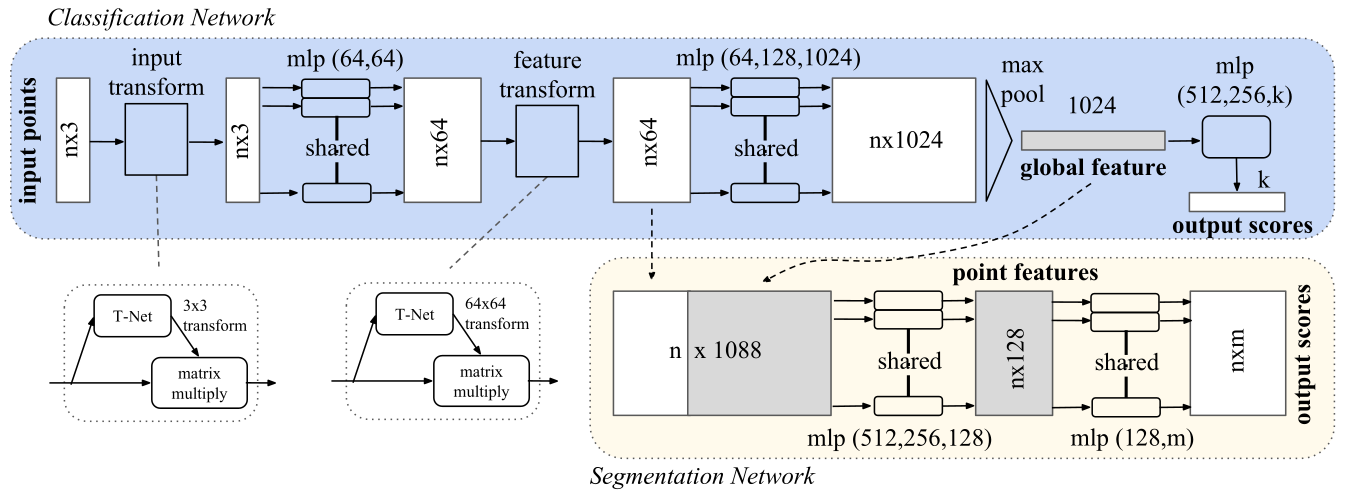
In 2023, I have implemented and integrated PointNet into the Figura environment and trained models for different non-defective anatomical regions, for instance the proximal and distal femur and the glenoid region of the scapula. At the 2025 International Society for Technology in Arthroplasty (ISTA) Conference I had the opportunity to present my work on the application of PointNet in the orthopedic field, titled:
Accuracy Evaluation of the PointNet Deep Learning Model Trained on Synthetic Data for the Segmentation of the Articular Surface and Intercondylar Fossa of the Healthy Femur
The aim of the work was to evaluate the performance of a segmentation model trained on synthetic point clouds representing the distal epiphysis of the femur. The articular surface and the intercondylar fossa were the two regions of interest taken into consideration.
The manual labelling of point clouds is a very time consuming tasks, thus the training and validation datasets were automatically annotated leveraging a statistical shape model (SSM) of the healthy femur.
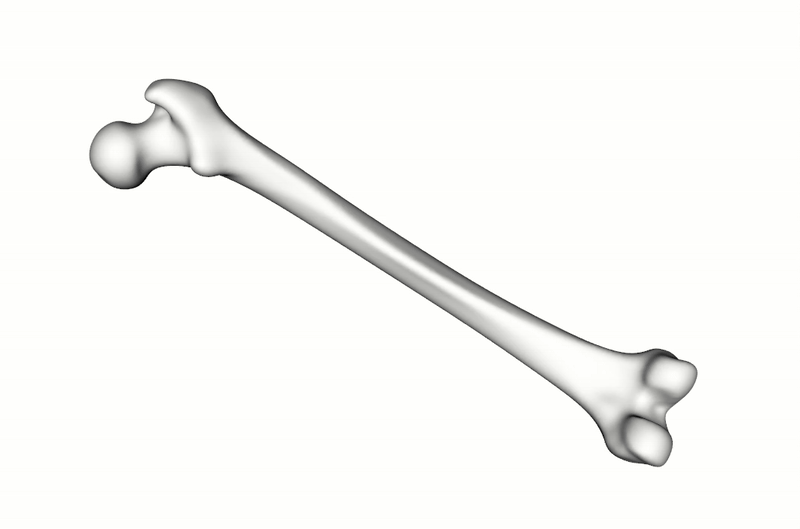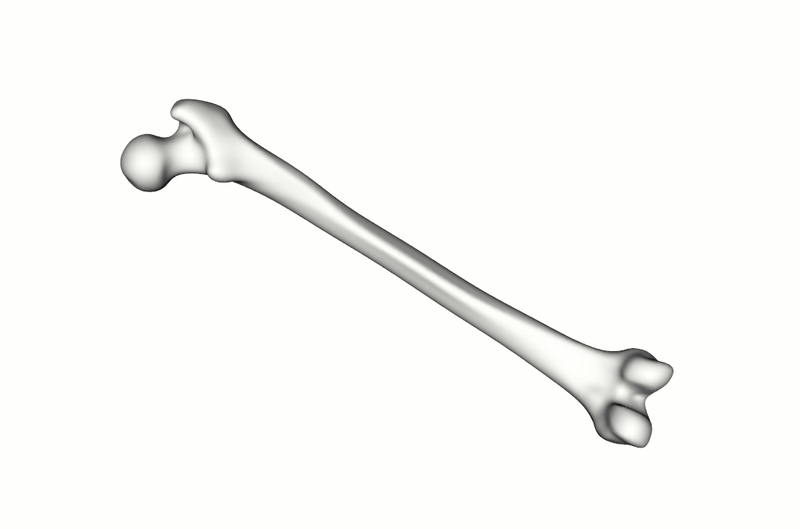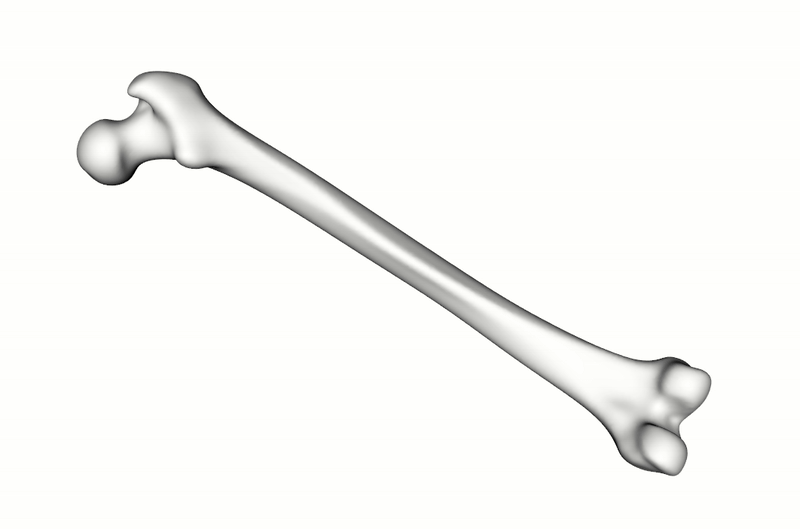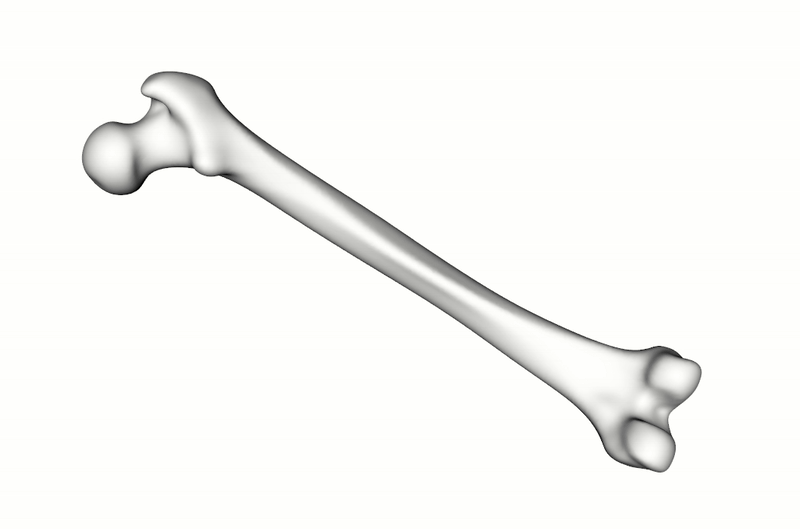
A total of 900 synthetic point clouds (800 for training, 100 for validation) were generated from the SSM which was built using 220 healthy adult femurs. As previously mentioned, the labels for the articular surface and intercondylar fossa were defined just once on the SSM reference and then automatically propagated to all synthetic instances using the SSM inherent point correspondences. To make the model more robust to orientation changes, the training data was augmented with random rotations.
The PointNet architecture was implemented in TensorFlow 2.13 using an exponential decay learning rate. Distributed training was performed across two NVIDIA RTX A2000 GPUs (CUDA 11.8) for 400 epochs with a batch size of 16.
Performance was then evaluated on a test set of 20 real, healthy femurs that were completely distinct from those used to build the SSM. Prior to processing, both the synthetic and real point clouds were truncated to isolate the distal region and uniformly downsampled to 8,192 points.
The model achieved a test accuracy of 0.95, test loss of 2.16, mean IoU of 0.89, and mean Dice coefficient of 0.94. The IoU values obtained for each class were 0.94 for background, 0.87 for the articular surface, and 0.85 for the intercondylar fossa, with corresponding Dice coefficients of 0.97, 0.93, and 0.92. The figure below illustrates the predicted segmentation results on a representative distal femur.
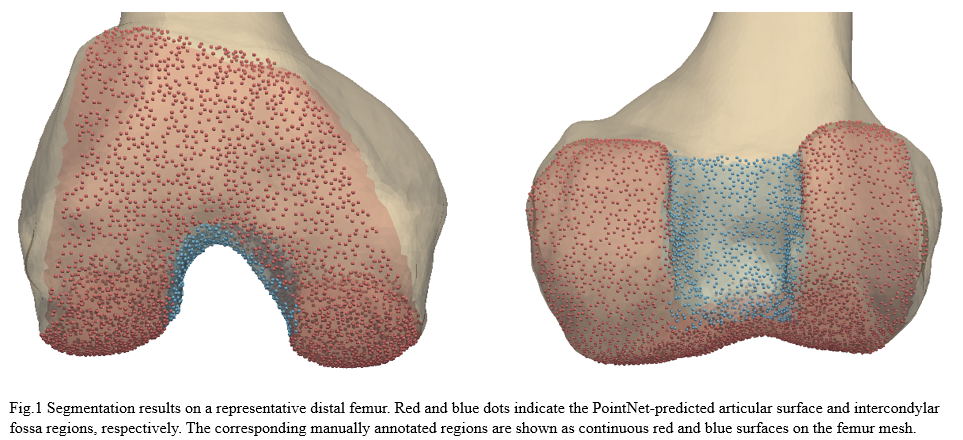
The results suggested that a PointNet model trained exclusively on synthetically generated data can generalize effectively to real anatomical point clouds, supporting the feasibility of leveraging statistical shape models for the generation of scalable and reproducible training datasets. This approach may reduce reliance on manual labeling while preserving good segmentation performance.
References
- Qi, C. R., Su, H., Mo, K., & Guibas, L. J. (2017). Pointnet: Deep learning on point sets for 3d classification and segmentation. Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 652–660.